Instrument driver
Once srsinst.rga is installed to your computer, you can use it to control and
acquire data from SRS RGAs in various Python environments,
such as Python interpreter console, Jupyter notebook,
context-sensitive editors, or just to write Python scripts.
Here we use the Python interpreter in interactive mode
to show how to use RGA100 class
to communicate with an RGA.
When you start Python from the command line, the ‘>>>’ prompt is waiting for your input.
C:\rga>python
C:\PyPI\rga>C:\Users\ckim\AppData\Local\Programs\Python\Python38\python.exe
Python 3.8.3 (tags/v3.8.3:6f8c832, May 13 2020, 22:37:02) [MSC v.1924 64 bit (AMD64)] on win32
Type "help", "copyright", "credits" or "license" for more information.
>>>
Connecting to an RGA
From the prompt, import RGA100 class
and connect to an RGA.
>>> from srsinst.rga import RGA100
>>> rga1 = RGA100('serial', 'COM3', 28800) # This is for a Windows computer
>>> rga1.check_id()
('SRSRGA200', '19161', '0.24')
In the case shown above, the RGA is connected to the serial port, COM3 on a Windows computer. Note that RGA 100 series only connects with the baud rate of 28800.
The check_id() method
reads the identification string from the RGA and
adjust the Scans component
depending on the highest mass.
Note that the serial port notation with a Linux computer is different from that of a Windows computer.
>>> rga2 = RGA100('serial', /dev/ttyUSB0', 28800) # for Linux serial communication
>>>
If your RGA is equipped with the RGA ethernet adapter (REA), it can be connected over Ethernet.
>>> rga3 = RGA100('tcpip', '192.168.1.100', 'admin',' admin')
>>>
You have to know the IP address, the user name and the password of the REA, to connect to an REA equipped RGA.
A general usage of RGA100
class can be simplified as:
Create an instance of RGA100 class
Connect to an RGA
Use it
Disconnect from it
>>>
>>> rga3 = RGA100()
>>> rga3.connect('tcpip','192.168.1.100','admin','admin')
>>> rga3.check_id()
('SRSRGA200', '19161', '0.24')
>>> rga3.disconnect()
>>>
Setting up ionizer
Let’s take a look into the ionizer component.
>>> pp.pprint( rga1.ionizer.dir )
{ 'components': {},
'commands': { 'electron_energy': ('RgaIntCommand', 'EE'),
'ion_energy': ('RgaIonEnergyCommand', 'IE'),
'focus_voltage': ('RgaIntCommand', 'VF'),
'emission_current': ('RgaFloatCommand', 'FL')},
'methods': ['get_parameters', 'set_parameters']}
>>>
It contains commands and methods to configure the ionizer of the RGA. Commands are defined using the Python descriptor to encapsulate raw RGA remote commands.
Each command item is defined as:
‘command name’: (‘Command class name’, the raw ‘RGA remote command’ that can be found in the RGA manual).
For example, the command electron_energy is defined using
RgaIntCommand
and it encapsulate the RGA remote command ‘EE’.
You can configure the ionizer parameters in various ways.
>>> rga1.ionizer.get_parameters()
(70, 12, 90) # tuple of (electron energy, ion energy, focus voltage)
>>> rga1.ionizer.electron_energy
70
>>> rga1.ionizer.electron_energy = 69
>>> rga1.ionizer.electron_energy
69
>>> rga1.ionizer.ion_energy
12
>>> rga1.ionizer.ion_energy = 8
>>> rga1.ionizer.ion_energy
8
>>> rga1.ionizer.focus_voltage
90
>>> rga1.ionizer.focus_voltage = 89
>>> rga1.ionizer.focus_voltage
89
>>> rga1.ionizer.get_parameters()
(69, 8, 89)
>>> rga1.ionizer.set_parameters() # set to default
0
>>> rga1.ionizer.get_parameters()
(70, 12, 90)
>>>
By defining a command as Python descriptor, it can be used as an attribute
instead of using raw communication functions
in RGA100 class with RGA raw commands.
>>> rga1.query_int('EE?') # equivalent to 'rga1.ionizer.electron_energy'
70
>>> rga1.query_int('EE69') # equivalent to 'rga1.ionizer.electron_energy = 69'
1
>>> rga1.query_int('EE?')
69
>>>
Turning filament on/off
You can turn the filament on or off, by adjusting the emission current in the ionizer component.
>>> rga1.ionizer.emission_current
0.3852
>>> rga1.ionizer.emission_current = 1.0
>>> rga1.ionizer.emission_current
1.0065
There is also the dedicated filament
component.
>>> pp.pprint( rga1.filament.dir )
{ 'components': {},
'commands': {},
'methods': ['turn_on', 'turn_off', 'start_degas']}
>>>
>>> print( rga1.filament.turn_on.__doc__ )
Turn on filament to the target emission current
Parameters
-----------
target_emission_current : int, optional
Default is 1.0 mA
Returns
--------
error_status : int
Error status byte
>>>
>>> rga1.ionizer.emission_current
0.0
>>> rga1.filament.turn_on()
0
>>> rga1.ionizer.emission_current
1.0076
>>> rga1.filament.turn_off()
0
>>>
Setting up detector
You can select the Faraday cup detector by setting CEM voltage to 0, and select Channel electron multiplier (CEM) detector and CEM voltage to a positive value.
>>> pp.pprint( rga1.cem.dir )
{ 'components': {},
'commands': { 'voltage': ('RgaIntCommand', 'HV'),
'stored_voltage': ('FloatNSCommand', 'MV'),
'stored_gain': ('RgaStoredCEMGainCommand', 'MG')},
'methods': ['turn_on', 'turn_off']}
>>> print( rga1.cem.turn_on.__doc__ )
Set CEM HV to the stored CEM voltage
>>> rga1.cem.stored_voltage
1043.0
>>> rga1.cem.voltage
0
>>> rga1.cem.turn_on()
>>> rga1.cem.voltage
1035
>>> rga1.cem.voltage = 0
>>> rga1.cem.voltage
0
Setting up a scan
Getting mass spectra is the core task of an RGA. All the functionality of acquiring
mass spectra resides in Scans class.
Let’s take a look what are available with the instance of
Scans class with the dir attribute.
>>> pp.pprint( rga1.scan.dir )
{ 'components': {},
'commands': { 'initial_mass': ('IntNSCommand', 'MI'),
'final_mass': ('IntNSCommand', 'MF'),
'speed': ('IntNSCommand', 'NF'),
'resolution': ('IntNSCommand', 'SA'),
'total_points_analog': ('IntGetCommand', 'AP'),
'total_points_histogram': ('IntGetCommand', 'HP')},
'methods': [ 'set_callbacks',
'set_data_callback_period',
'get_data_callback_period',
'get_max_mass',
'get_parameters',
'set_parameters',
'read_long',
'get_mass_axis',
'get_analog_scan',
'get_histogram_scan',
'get_multiple_mass_scan',
'get_single_mass_scan',
'set_mass_lock',
'get_partial_pressure_corrected_spectrum',
'get_peak_from_analog_scan']}
>>>
To set up a scan, we have to specify the initial mass, final mass, scan speed, and resolution (steps per AMU).
>>> rga1.scan.get_parameters()
(2, 50, 3, 20)
>>> rga1.scan.initial_mass = 1
>>> rga1.scan.final_mass = 65
>>> rga1.scan.speed = 4
>>> rga1.scan.resolution = 10
>>> rga1.scan.get_parameters()
(1, 65, 4, 10)
>>> rga1.scan.set_parameters(10, 50, 3, 20)
>>> rga1.scan.get_parameters()
(2, 50, 3, 20)
>>>
Acquiring a histogram scan
>>> rga1.scan.set_parameters(10, 50, 3, 20)
>>> histogram_mass_axis = rga1.scan.get_mass_axis(for_analog_scan=False)
>>> histogram_mass_axis
array([10., 11., 12., 13., 14., 15., 16., 17., 18., 19., 20., 21., 22.,
23., 24., 25., 26., 27., 28., 29., 30., 31., 32., 33., 34., 35.,
36., 37., 38., 39., 40., 41., 42., 43., 44., 45., 46., 47., 48.,
49., 50.])
>>>
>>> histogram_spectrum = rga1.scan.get_histogram_scan()
>>> histogram_spectrum
array([ 211., 175., 56., 24., 249., 129., 213., 303., 639.,
533., 217., 191., 206., 179., 256., 116., -116., 222.,
343., 240., 206., 249., 347., 483., 20., -65., 104.,
249., 179., 307., 239., 245., 347., 262., 312., 226.,
307., 271., 468., 226., 201.])
Acquiring a analog scan
>>> rga1.scan.set_parameters(1, 50, 3, 20)
>>> mass_axis = rga1.scan.get_mass_axis(for_analog_scan=True)
>>>
>>> spectrum = rga1.scan.get_analog_scan()
>>> spectrum_in_torr = rga1.scan.get_partial_pressure_corrected_spectrum(spectrum)
Saving a spectrum to a file
>>> with open('spectrum.dat', 'w') as f:
... for x, y in zip(mass_axis, spectrum_in_torr):
... f.write('{:.2f} {:.4e}\n'.format(x, y))
Plot a spectrum with matplotlib
>>> import matplotlib.pyplot as plt
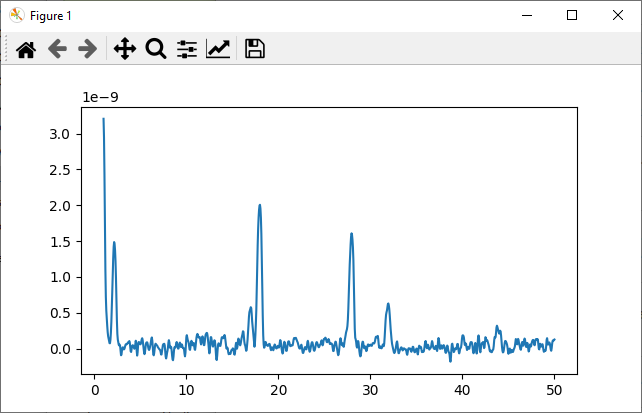
>>> plt.plot(mass_axis, spectrum_in_torr)
>>> plt.show()
It will bring up a plot showing an analog scan spectrum.
As a summary, the following script is put together to get an analog scan plot from the beginning.
import matplotlib.pyplot as plt
from srsinst.rga import RGA100
rga1 = RGA100('serial','COM3', 28800)
rga1.filament.turn_on()
rga1.cem.turn_off()
rga1.scan.set_parameters(1, 50, 3, 20) # (initial mass, final mass, scan speed, resolution)
mass_axis = rga1.scan.get_mass_axis(True) # True for analog scan, False for histogram scan
spectrum = rga1.scan.get_analog_scan()
spectrum_in_torr = rga1.scan.get_partial_pressure_corrected_spectrum(spectrum)
rga1.disconnect())
plt.plot(mass_axis, spectrum_in_torr)
plt.show()